|
|
|
|||||

Biomedical Applications of Vibrational Spectroscopy - Hyperspectral Imaging - Chemometrics
|
||||||
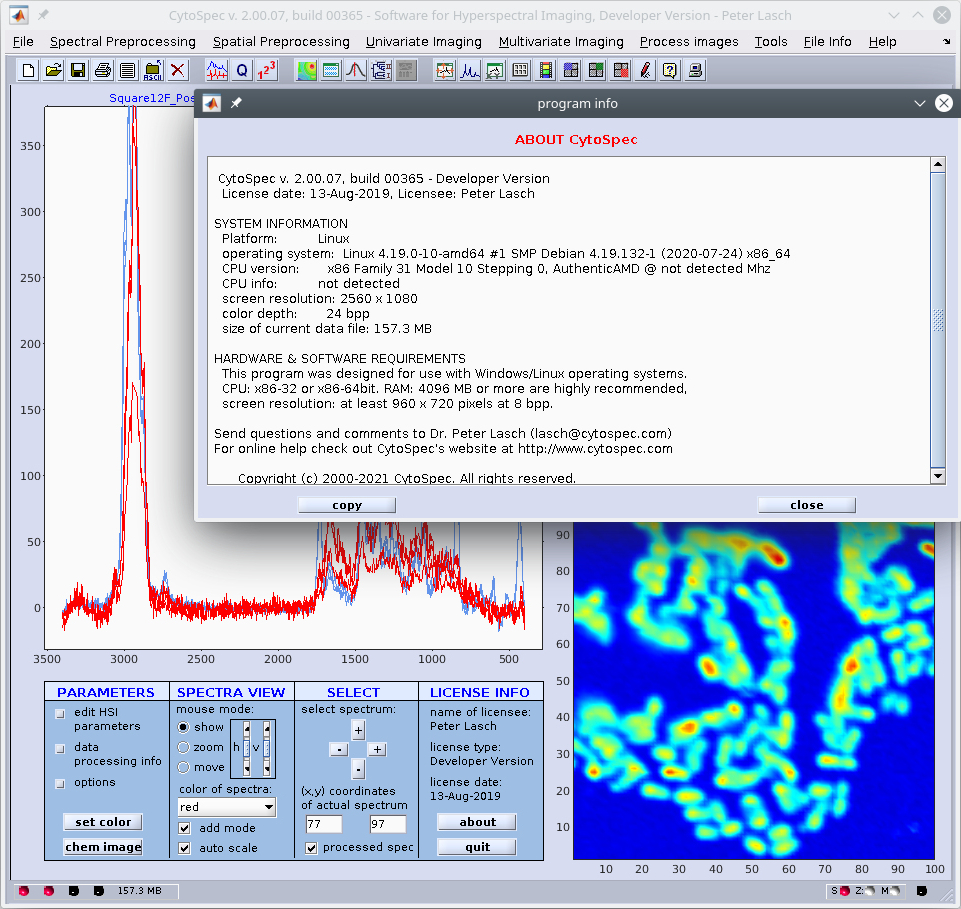
CytoSpec for LINUX
|
Links:
|
 The Mathworks
The Mathworks  here. Note that a license key must be ordered separately. To register
send an e-mail with your request to
service@cytospec.com. here. Note that a license key must be ordered separately. To register
send an e-mail with your request to
service@cytospec.com. |
|
[
GENERAL |
FILE |
SPECTRAL PREPROCESSING |
SPATIAL PREPROCESSING |
UNIVARIATE IMAGING |
MULTIVARIATE IMAGING |
TOOLS |
FILE INFO |
GLOSSARY |
CONTACT: info@cytospec.com |
PUBLISHER DETAILS |
PRIVACY POLICY ]
Copyright (c) 2000-2026 CytoSpec. All rights reserved.
FILE INFO |
Copyright (c) 2000-2026 CytoSpec. All rights reserved.